
The throughput does increase linearly with the number of allocated CPUs from 32 to 50.
#Samtools threads full
As for alternative A we testedĭifferent numbers of CPUs - from a full allocation of 50 CPUs down toģ2 CPUs to account for the fact that all three tools don’t operate at Some initial experiments showed that 42GB of memory were (ubam) and includes some additional steps but for the purposes of this tutorialįastp -i $read1 -I $read2 \ -stdout -thread 2 \ -j " $.logįor an input file with 201,726,334 read pairs we would expect a runtime ofĪbout 2.5h given the rate limits of ~22,000 pairs/sec in theĪlignment step. Note that the GATK best practices pipeline starts from an unaligned BAM file Whether sorting or not, it is important to not write uncompressed SAMįiles to disk since conversion to BAM is not computationally intensive and BAM Similar to each other and within the range of good parallel efficiency (70~80% Of each of the three components should be Unmodified or sorted BAM output from FASTQ files are presented.įor efficiency trimming, alignment, and converting (or sorting) to BAM format are runĬoncurrently with data flowing through linux pipes to avoid unnecessary IO. Therefore, in this Chapter benchmarks and scripts for either However, in other cases a sorted BAM file prior to marking duplicates The issue that I encountered is that the speed of 'calmd' is incredibly slow The jobs have already run 12 hours, and only 600MB BAM with MD tag are generated. The size of original BAM is around 50Gb (whole genome sequence by using pacbio HIFI reads). Output so the most efficient pipeline would not write sorted BAM files from theĪligner. I am using 'samtools calmd' to add MD tag back to BAM file. Queryname-grouped input as generated by the aligner and produce sorted BAM Bgzip-compressed and tabix-indexed file with annotations.
Add or remove annotations.-a, -annotations file. The option is currently used only for the compression of the output stream, only when -output-type is b or z. GATK’s duplicate marking tools perform more efficiently with Use multithreading with INT worker threads. Duplicate marking itself is discussed in Chapterģ. Thread support first arrived in version 0.1.19 (March 2013), which enabled them for sorting and BAM file writing in the view command. 8.3 Optimized script for VariantRecalibratorĭata preprocessing includes read trimming, alignment, sorting by coordinate,Īnd marking duplicates.6 GenomicsDBImport (replaces CombineGVCFs).4.3 Optimized script for BaseRecalibrator.3.2.3 Performance comparing between queryname-grouped and coordinate-sorted inputs.
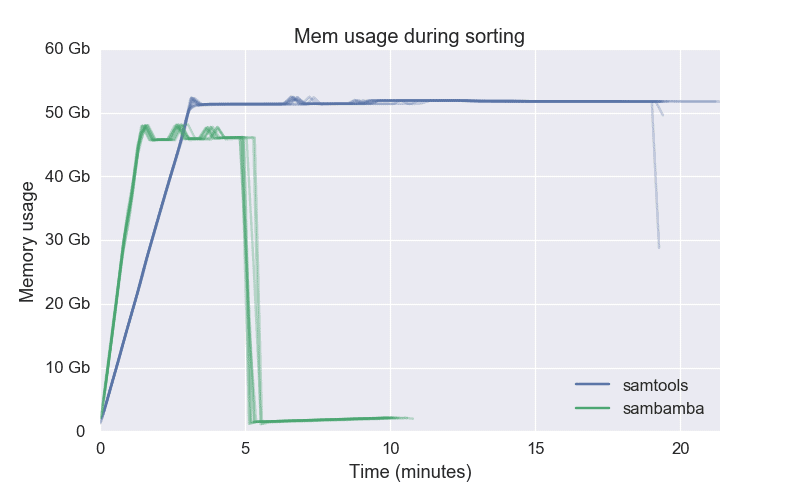
3.2.1 Queryname-grouped input data (as generated by the aligner).1.4.2 Alternative B: BAM output without coordinate sorting.1.4.1 Alternative A: producing sorted BAM output.1.3.4 Alternative B: Queryname-grouped (unsorted) BAM.The approximate peak memory usage (MaxVMSize) recorded for SAMtools 1.3. There was little variation in runtimes between either version at any thread count: the average RSD was 0.39 for SAMtools 1.3.1, and 0.40 for SAMtools OpenMP. 1.3.3 Alternative A: Sorting by coordinate At (ge 8) threads, SAMtools OpenMP outperformed SAMtools 1.3.1 by approximately a factor of 2.Some other workflow schedulers allow you to quote the entire command line, but a glance at the srun man page seems to indicate that this wouldn’t work.

#Samtools threads how to
I don’t use Slurm so I’m not entirely sure how to correct the command line.
